One of the most rewarding parts of building SimAtomic is seeing it used in real research, especially when the workflow ends up in a peer-reviewed paper.
Recently, SimAtomic had the opportunity to collaborate with Florida State University (FSU) Research Foundation to provide molecular dynamics support for an ACS Omega study by Helene Tigro and Christopher Solís, “Desensitization of the Cardiac Troponin Complex by TnI Phosphorylation and Epigallocatechin-3-Gallate.”
I am not a co-author, SimAtomic's contribution was computational MD support.

Desensitization of the Cardiac Troponin Complex by TnI Phosphorylation and Epigallocatechin-3-Gallate, ACS Omega (January 13, 2026)
Background
Cardiac contraction depends on thin-filament sensitivity to Ca²⁺, which is regulated by the troponin complex (TnC–TnI). In some heart diseases this sensitivity is abnormally high, impairing relaxation, so Ca²⁺ desensitization is a potential therapeutic strategy. The authors tested two desensitizing interventions: (1) phosphorylation of TnI at Ser23/24 and (2) the small molecule EGCG. Here, I focus on Ser23/24 phosphorylation—the part that included MD simulations. They paired in vitro motility assays with structural modeling (AlphaFold 3 and MD) to connect functional changes to a mechanistic structural explanation.
By running MD on AlphaFold 3-generated wild-type (WT) troponin structures (n = 11) and measuring hydrogen-bond distances between TnI and TnC, they showed that the predicted interface is dynamically plausible. The transient, flickering hydrogen-bond connections also made it more plausible that phosphorylation at Ser23/24 can regulate the complex by shifting this contact network.
"AlphaFold is great for hypothesis generation, but we must validate these statistical guesses with physics, and this is why MD simulations are now more relevant than ever."
Computational Workflow
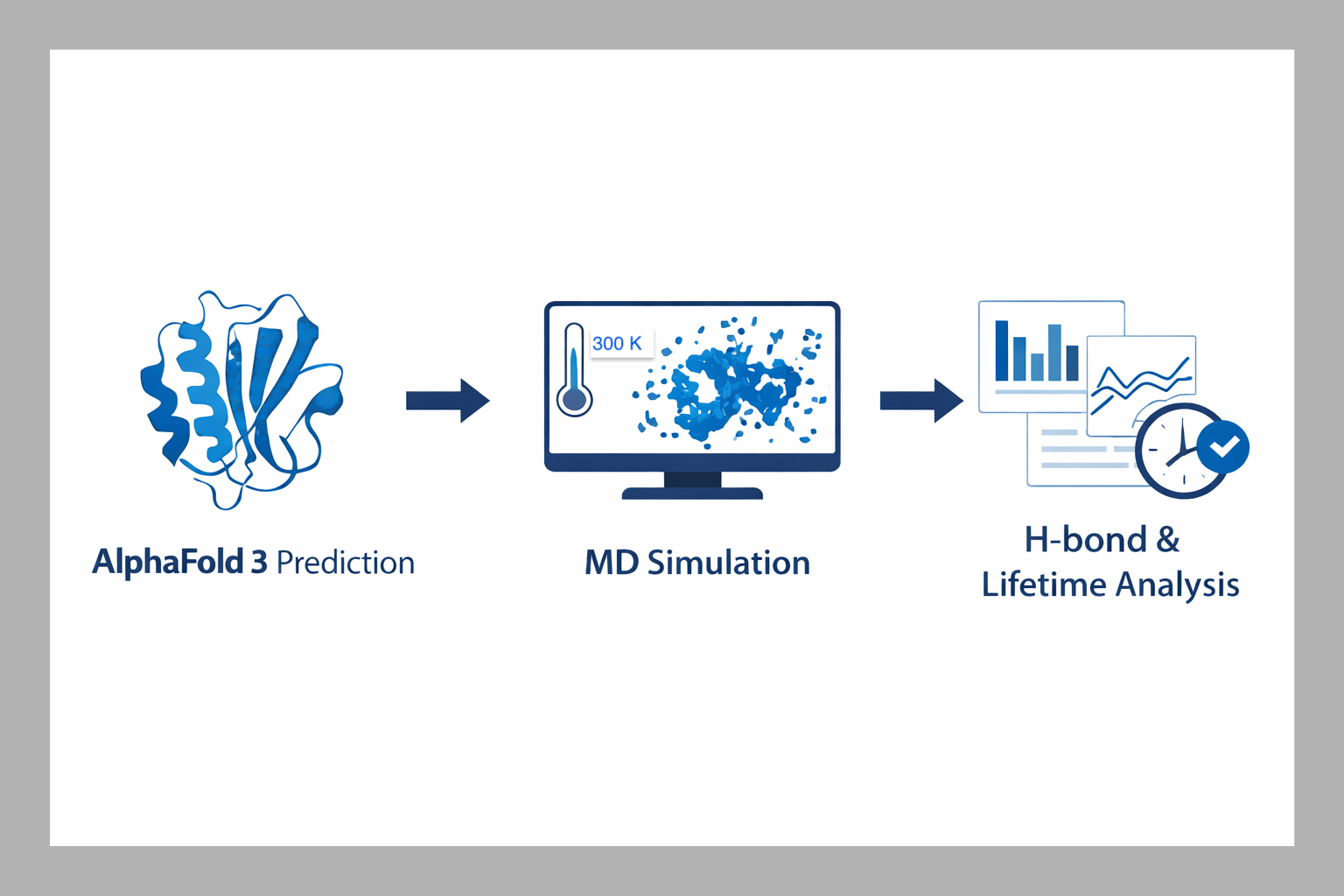
Overview of the computational workflow used in the FSU study
1. Start from AlphaFold 3 structures
To understand how phosphorylation of TnI at Ser23/24 alters the troponin complex, the authors used AlphaFold 3 to generate 3D structural models (n = 11) in two states, wild type (WT) and Ser23/24-phosphorylated, and compared how the complex might change at the atomic level. To reduce “model hallucination” risk, they applied quality filters:
- pTM > 0.7 (global confidence)
- pIDDT > 0.5 for the S23/24 region
so they only analyzed models where the phosphorylation site neighborhood had reasonable confidence. Because AlphaFold 3 outputs static snapshots, they then ran molecular dynamics (MD) simulations on SimAtomic to check whether the TnI–TnC contacts suggested by the AlphaFold models were dynamically plausible over time, rather than artifacts of a single predicted conformation.
2. Trim to the essentials
Before simulating, they simplified the system to focus on the regions of interest. They removed:
- the TnT subunit
- TnI residues 1–15
- TnI residues 151–210
3. Run MD using SimAtomic
MD simulations were performed using SimAtomic (online platform) under a standard, neutralizing-condition preset. Each trajectory was run for 1 ns total, with frames saved every 0.04 ps. Simulations were performed at pH 7.2 and 15 mM NaCl.
- Total simulation length: 1 ns
- Frame spacing: 0.04 ps per frame
- Time step: 2 fs per step
- Force field / water model: AMBER19SB with TIP3P water
- Boundary conditions: Periodic boundary conditions
- Ensemble: NVT (constant volume; no barostat/pressure coupling)
- Temperature control: Langevin dynamics with 300 K
- Electrostatics: Particle Mesh Ewald (PME) with a nonbonded cutoff distance
4. Analyze trajectories to quantify key H-bond contacts
The authors exported the simulation outputs (structures and trajectories) and then used ChimeraX to track, frame by frame, how close specific parts of troponin stayed—focusing on hydrogen-bond–like contacts between the N-terminal of TnI (residues 1–34) and the N-lobe of TnC. They saved these measurements to a text file and analyzed them in R to make clean time-series plots and summary histograms.
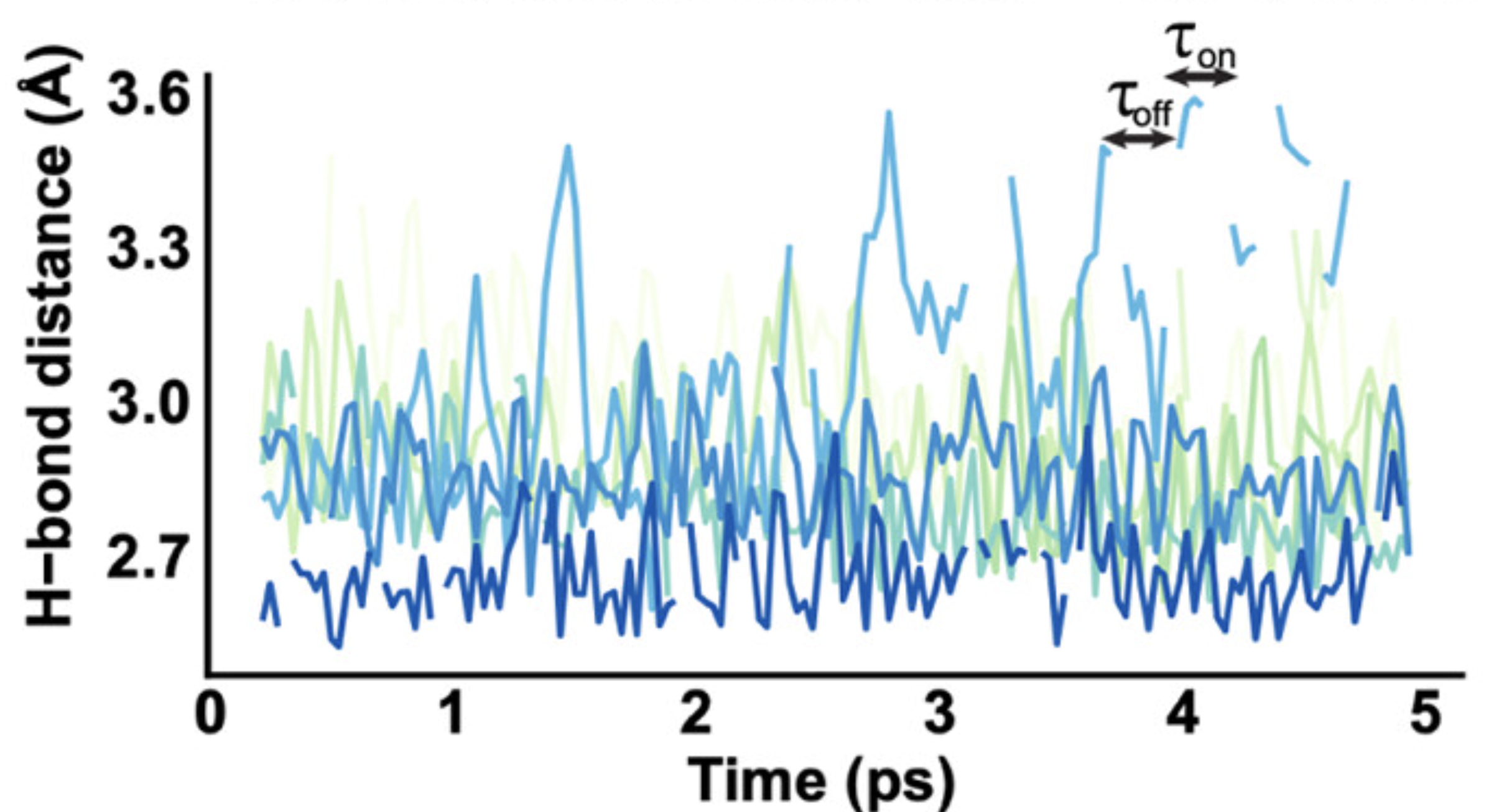
Figure 3F from Solís et al., ACS Omega (2025). Hydrogen bond distance traces from MD simulations of wild-type troponin complexes.
What these plots show: The time-series plots display hydrogen bond distances between TnI and TnC residues over the course of 5 ps MD simulations. Each trace represents a different hydrogen bond contact, with distances plotted on the y-axis (typically in the 2–4 Å range for hydrogen bonds) and simulation time on the x-axis. The plots revealed that these contacts were highly dynamic. They repeatedly formed and broke throughout the simulation, rather than remaining static. This flickering behavior supported the idea that the TnI–TnC interface is flexible and can be modulated by phosphorylation or small molecules like EGCG.
This analysis turned a static idea ("these regions can touch") into a dynamic result ("they touch often, in a realistic way"), which strengthened the proposed mechanism for how phosphorylation (and EGCG) can tune Ca²⁺ sensitivity through troponin.
The Value of a Streamlined Workflow
If you've ever tried to go from "I have a structure" to "I have a figure and a quantitative story," you know how many tiny steps can derail momentum. This paper is a great example of keeping the workflow moving: generate hypotheses with structure prediction and docking, then use short MD to test whether key interactions are dynamic and plausible.
What we learned
This collaboration highlighted that the bottleneck in ensemble MD is often post-processing: running many simulations in parallel and turning trajectories into consistent, comparable metrics without file conversions and custom scripts.
In response, SimAtomic added built-in hydrogen-bond occupancy analysis (summary contact stability) and an API endpoint to submit multiple jobs in parallel and download standardized results bundles directly to a laptop. We also standardized outputs and run metadata to make it easier to aggregate and reproduce analyses across models and conditions.
Want to Run a Similar Workflow?
If you're working on protein–protein interfaces, allostery, or ligand effects, then this kind of "fast MD + clear analysis" can be the difference between a plausible model and a publishable story.
If you’re preparing a manuscript or stress-testing a structural hypothesis, SimAtomic can help you:
- go from structure → MD trajectory quickly,
- export standard files for your preferred analysis toolchain, and
- keep the workflow reproducible end-to-end.
And if you're at a university lab like our collaborators at Florida State University, I'd love to hear what you're working on.
Reference
Desensitization of the Cardiac Troponin Complex by TnI Phosphorylation and Epigallocatechin-3-Gallate, ACS Omega (collection date Jan 13, 2026; originally published Dec 20, 2025).